What we propose
}Bridging the Translational Gap
The challenge
}Why Promising Drugs Still Fail
A lot of promising drugs fail in clinical trials, leaving patients waiting for treatments they desperately need.
High Failure Rates
The overall success rate of a drug passing through all clinical trials is 8.4%. Depending on the therapeutic area the range is from 3.9% for infectious disease to 34.1% for blood borne diseases.
Lack of Efficacy
Phase 2 trials that test if the drug is effective, have the lowest probability of success, ranging from 27.4% for respiratory diseases to 52.7% for ophthalmology.
Unclear MoA
Lack of Efficacy results from poorly understood disease biology. How does the target to which the drug binds to modulate the disease (Mechanism of Action - MoA)?
How we do it
}We Tackle the Root Cause.
Causal simulation. Biomarker trajectory modeling. Mechanism-aware target evaluation.
Built for you, To decode biology.
- From static snapshots to dynamic insights
- From prediction patterns to prognostic simulation
- Simulate relative system behavior under different target perturbations to compare mechanistic hypotheses
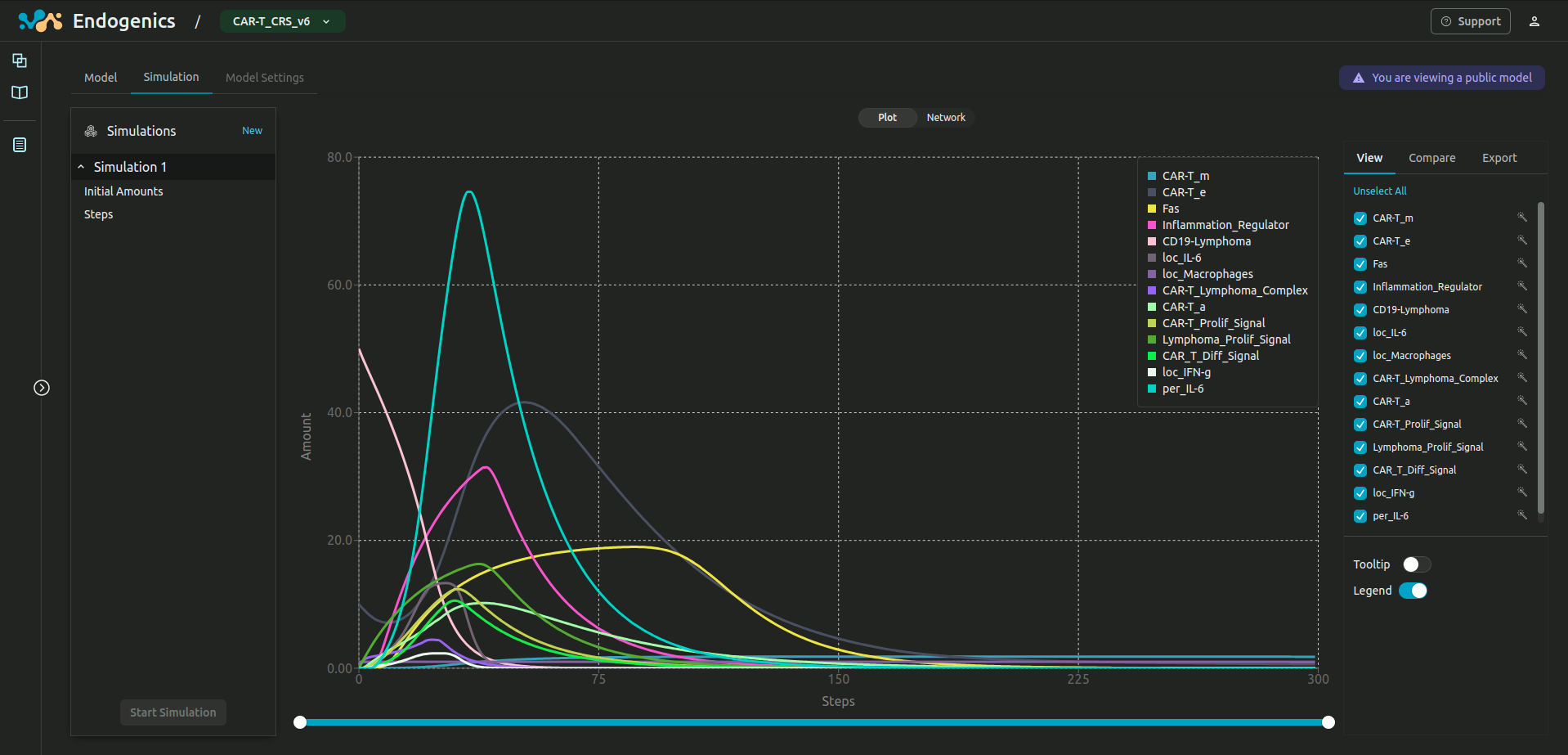
EndoGenics™
BetaA mechanism-based modelling and simulation platform. Developed from first principles.
See more
EndoClinics ™
Coming soonA customized user interface for insights of potential clinical outcomes on the basis of in-house and public research data on COVID.
Our advancements
}News and Insights
Our Collaborators and Supporters
We are just a click away to discuss your project





